2026 Excellence in Environmental Engineering and Science® Awards Competition Winner
Honor Award - University Research
AI-Driven Machine Learning Frameworks for Optimizing Biomimetic Algal Biofuel Production
Entrant: Purdue University
Engineer in Charge: Zhi Zhou, Ph.D., P.E., BCEE
Location: West Lafayette, Indiana
Entrant Profile
The entrant, Dr. Zhi Zhou, served as the Principal Investigator and Lead Researcher for this multi-phase research project. He conceptualized the biomimetic approach of harnessing natural processes to disrupt algal cells without energy-intensive sonication or chemicals and pioneered the integration of artificial intelligence (AI) methods to address longstandingbottlenecks in algal biofuel production. In this capacity, Dr. Zhou oversaw experimental design, supervised data collection, and directed the development of AI-driven machine learning frameworks to optimize lipid productivity. This project represented a collaborative, multidisciplinary effort at Purdue University.
Dr. Zhou and his former Ph.D. student, Zhe Sun, developed a novel algal cell disruption technique that achieved lipid yields comparable to sonication while reducing energy requirements by 91.7%. His Ph.D. student, Amanda M. Lopez, led single-cell phenotypic analyses, developed unsupervised machine learning models (including PCA and K-means clustering), and created a random forest AI framework based on 38 cultivation variables and over 12,000 data points, predicting high lipid productivity with 95.2% accuracy. Undergraduate research assistant Yoonjung Choi supported algal cultivation experiments and metabolic data acquisition, contributing to a 165.2% relative increase in triacylglycerol accumulation. Additionally, collaborator and Ph.D. student Sean Savage provided expertise in data analysis, refining the predictive performance of ensemble decision tree algorithms.
Together, this team bridged environmental engineering and machine learning to advance the state-of-the-art in sustainable biofuel production. Their collective efforts yielded a patent, multiple peer-reviewed publications, and a comprehensive, data-driven framework for cost-effective, environmentally friendly renewable energy.
Project Description
Integrated approach
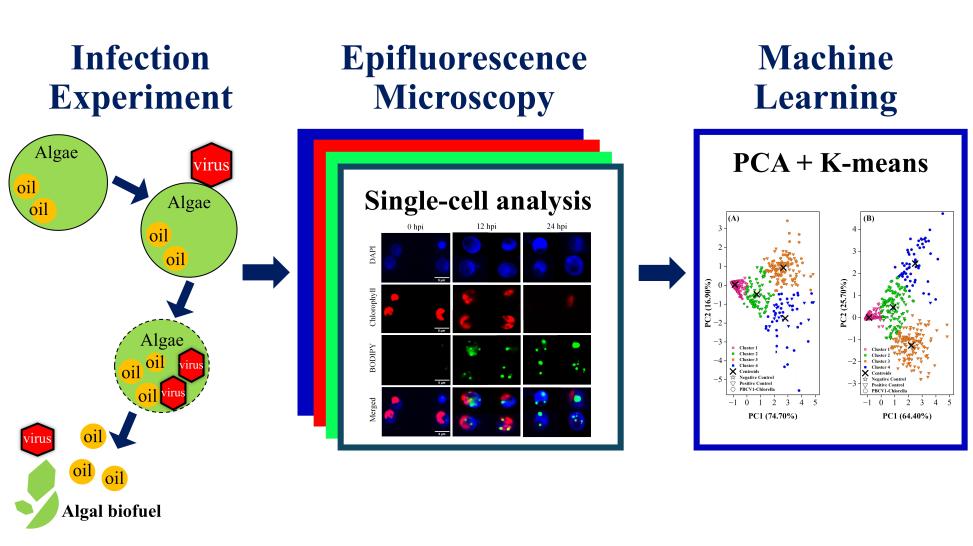
Algal biofuels offer a sustainable pathway to renewable energy and microalgae are capable of rapid growth, high lipid yields, and CO2 sequestration without competing for arable land or compromising environmental quality. However, commercialization is hindered by high production costs from energy-intensive harvesting, cell disruption, and lipid extraction (50–65% of total expenses) and the persistent trade-off between biomass productivity (BP) and lipid content (LC). This multi-phase research addresses these challenges by integrating nature-inspired biomimeticcell lysis with advanced AI/ML techniques. Spanning from a foundational patent on low-energy disruption to 2025–2026 studies on single-cell phenotyping, the project delivers transformative results of a 91.7% reduction in disruption energy, a 165.2% relative increase in lipid accumulation without stress induction, and a 95.2% accuracy rate inpredicting lipid productivity (LP) conditions. As shown in the following figure, this holistic and integrated AI-driven framework redefines algal biofuel production as efficient, scalable, and environmentally benign.
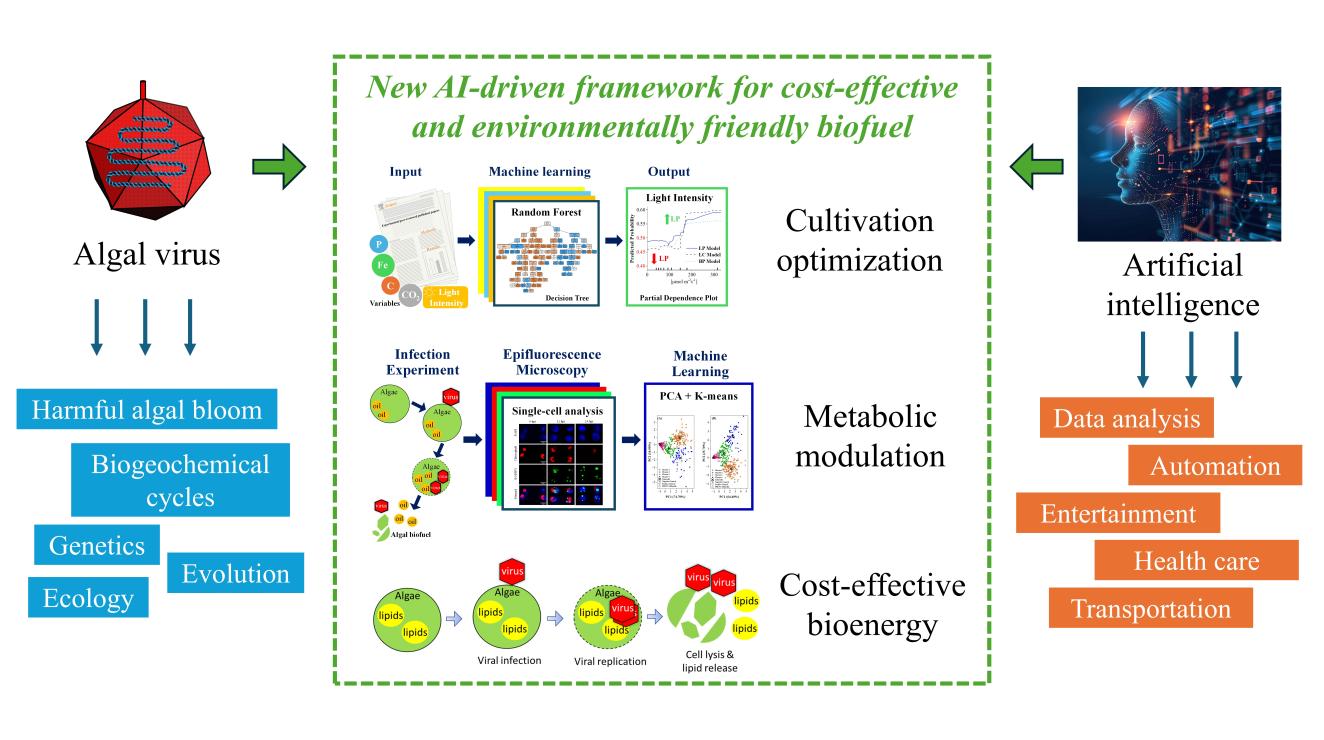
The project exemplifies an integrated approach that mitigates nutrient pollution and energy consumption across air, water, and land, ensuring solutions do not merely shift environmental burdens. While conventional methods rely on energy-intensive ultrasonication or chemical solvents—consuming substantial fossil-derived electricity and generating toxic waste—this biomimetic lysis technique leverages natural cellular processes under ambient conditions. The patented method achieves complete disruption of concentrated Chlorella biomass with minimal input, slashing energy use by 91.7% and eliminating chemical byproducts that contaminate soil or groundwater. Furthermore, high-resolution epifluorescence microscopy of over 4,000 cells confirms lipid enrichment under nutrient-replete conditions, preventing nutrient runoff and eutrophication risks. Finally, a Random Forest ML model optimizes 38 cultivation variables to reduce the operational footprint: requiring less water, minimal land, and enhanced CO2 fixation. This closed-loop system promotes positive cross-media benefits, aligning perfectly with the principles of sustainable bioenergy.
Quality
Quality is substantiated by rigorous methodology, reproducibility, and a distinguished record of a patent, publications, and awards. The core biomimetic lysis technology was granted U.S. Patent No. 11,149,290, highlighting its innovative nature and translational impact. This technology has garnered multiple awards and national recognition for its sustainable approach to algal biofuel production.
In 2022, the team advanced as Technology Bonus Prize Semifinalists (Great Lakes Region) in the U.S. Department of Energy (DOE) EnergyTech University Prize. Building on this momentum in 2023, the project earnedthe Audience Choice Award at the Purdue Innovates Startup Expo’s New Venture Challenge, secured Regional Champion status, and again reached the Technology Bonus Prize Semifinalist stage in the DOE’s Fossil Energy and Carbon Management category. Most recently, in 2024, theproject progressed as a National Finalist (Top 6.7%) in the DOE EnergyTech University Prize. These distinctions underscore the technology’s potential to transform carbon-negative biofuel processing and reflect its resonance within prestigious national energy innovation circles.
The project team further expanded this technique by integrating an AI/Machine Learning framework to optimize biomimetic production. This work is documented in prestigious journals, including Bioresource Technology (2025; lipid enrichment), Renewable Energy (2026; ML optimization), and Data in Brief (2025; dataset), adhering to the higheststandards of experimental validation and peer review.
The 2025 single-cell dataset captures multidimensional phenotypes, chlorophyll autofluorescence, and nuclearmorphology, reusable resource for ML training that uncovers cellular heterogeneity overlooked in bulk assays. The 2026 Random Forest model, trained on a curated database of over 12,000 data points, outperforms traditional optimization tools by effectively handling nonlinearity and data imbalance. Furthermore, the team’s engagement with over 100 bioenergy experts through the NSF Innovation Corps (I-Corps) program confirmed the industrial necessity of resolving metabolic trade-offs. These endorsements align with U.S. DOE priorities in biosystems design. Through reproducible protocols—including fixed microscopy exposures, calibrated ion chromatography, and ensemble ML aggregation—this research establishes a new benchmark for advancing the viability of algal biofuels.
Originality and innovation
Originality and innovation are central, combining new applications of existing knowledge with novel hybrids. While prior efforts emphasized nutrient starvation, salt stress, or genetic modification to boost lipids (oftensuppressing growth), this project introduces biomimetic lysis as a non-stress metabolic modulator, achieving TAG levels rivaling stress techniques without confounders. The 2025 study first revealed a 165.2% lipid surge in infected Chlorella via unsupervised ML (principal component analysis and K-means clustering on single-cell data), mapping asynchronous trajectories and decoupling size from fluorescence for true metabolic shifts. This marks the inaugural single-cell resolution of lytic-cycle phenomics in biofuel contexts. The 2026 innovation of the first random forest ensemble for LP prediction challenges nitrogen-centric paradigms by identifying exogenous iron and phosphorus as primary drivers via partial dependence plots. Unlike single decision trees or parametric response surface methodology (scaling poorly with variables), this non-parametric approach aggregates hundreds of trees, mitigating overfitting and inter-studyvariability for genus-wide generalizability. The open-access 2025 dataset further enables community-driven algorithm development, fostering interdisciplinary advances in computational biology.
Complexity
The complexity addressed is multifaceted, involving biological variability, nonlinear parameter interactions, and engineering scalability. Algal systems exhibit cell-to-cell heterogeneity, transient states during cycles, and confounding factors (e.g., nutrient stress mimicking desired effects). Traditional factorial experiments for 38 variableswould demand billions of trials, while the project team dissects these via high-throughput imaging, quantitative fluorescence extraction in ImageJ, and nutrient profiling to isolate mechanisms. ML handles imbalanced data, taxonomic nuances, and overfitting, while revealing subpopulations resilient to trade-offs. Inland cultivation adds constraints like water/land availability, amplified by climate variability, and the project team’s narrowed ranges and predictive accuracy directly mitigate these environmental challenges.
Contribution to environmental, social, and economic advancement
Finally, the project drives social and economic advancement by making algal biofuels competitive. Current algalcultivation costs stem from cultivation/extraction inefficiencies. The project team’s innovations promise 50-65% reductions via energy savings and yield boosts, enabling commercial viability. This supports energy security, job creation in biotechnology (especially rural areas via inland farms), and equitable access to clean fuels. Its environmental benefits are achieved through carbon-negative cycles via CO2uptake and wastewater integration, contributing to global climate goals and UN Sustainable Development Goals(affordable and clean energy, climate action). The patent, novel AI framework, and open datasets position this technology for broad adoption, accelerating the transition to renewables and delivering long-term societal benefits in sustainability and economic resilience.
Click images to enlarge in separate window.
 |
 |
|
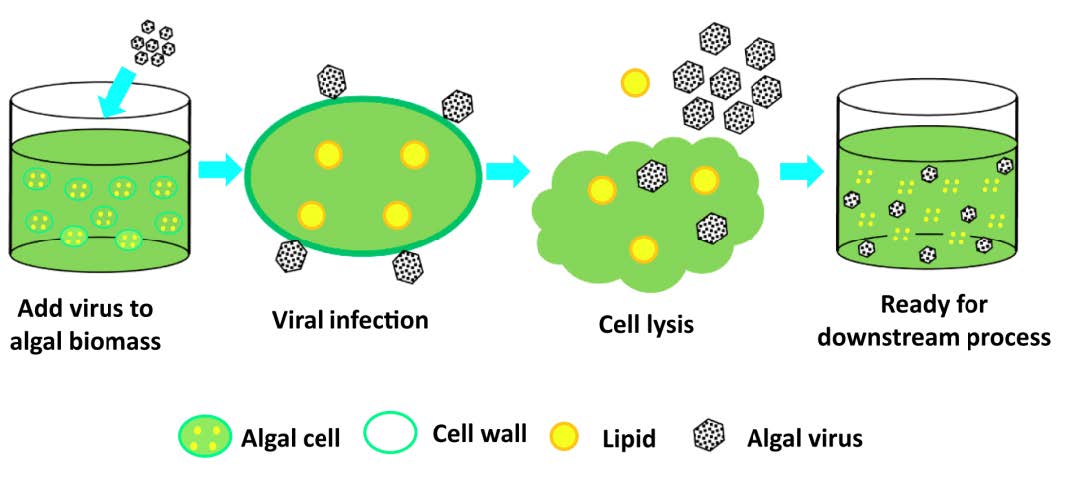
Figure 1. Nature-inspired virus-assisted algal cell disruption for cost-effective biofuel production
|
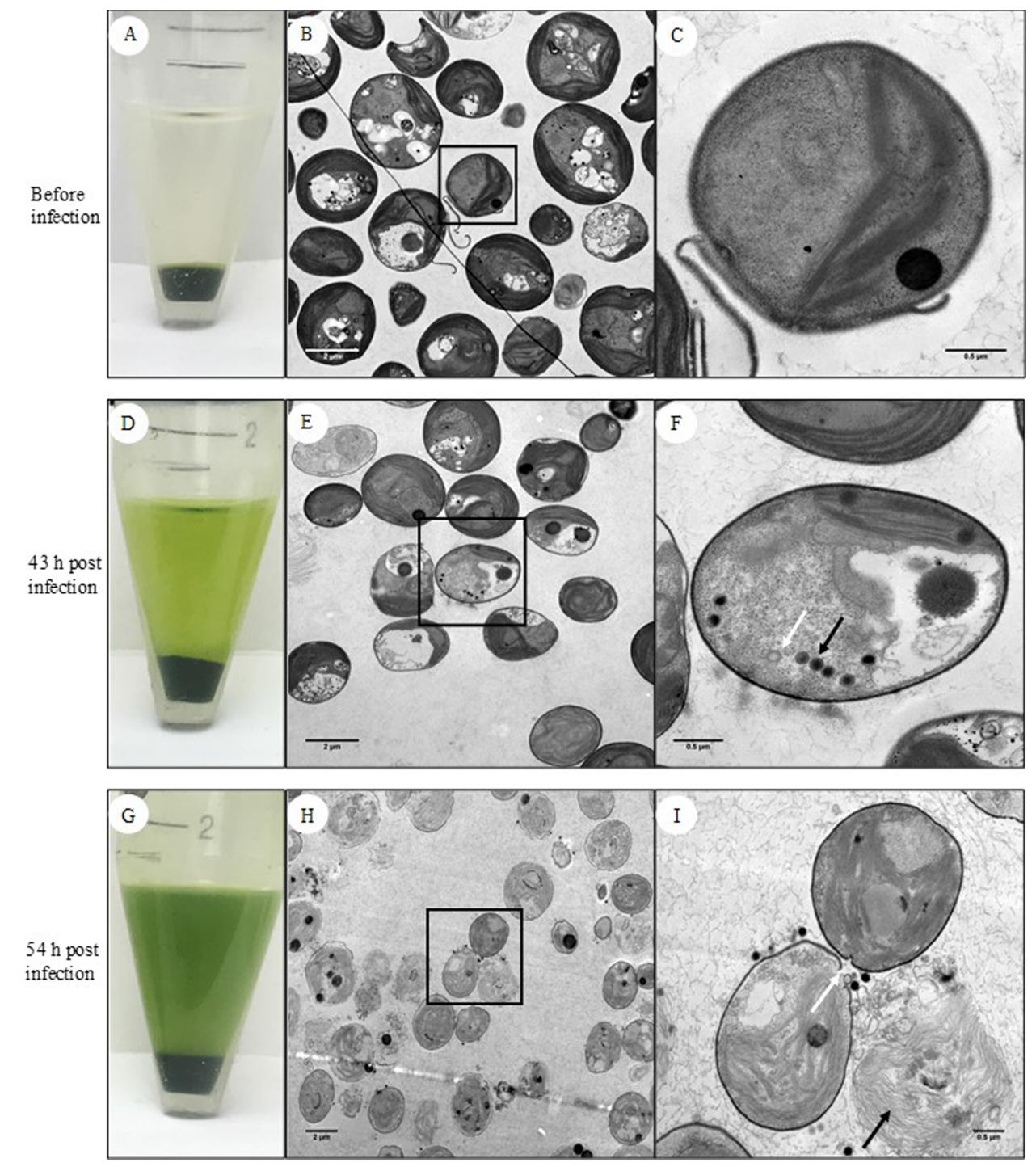
Figure. 2. Chlorella cells before and after viral infection. (A) Uninfected cells; (B) TEM picture of uninfected cells; (C) uninfected cell under high resolution; (D) Infected cells 43 h post infection; (E) TEM picture of infected cells 43 h post infection; (F) Infected cell 43 h post infection under high resolution showing assembled virus particles (black arrow) and empty virus capsids (white arrow); (G) Infected cells 54 h post infection; (H) TEM picture of infected cells 54 h post infection; (I) Infected cell 54 h post infection under high resolution showing highly deteriorated cell wall (black arrow) and empty virus capsid and degraded cell wall (white arrow).
|
 |
 |
|
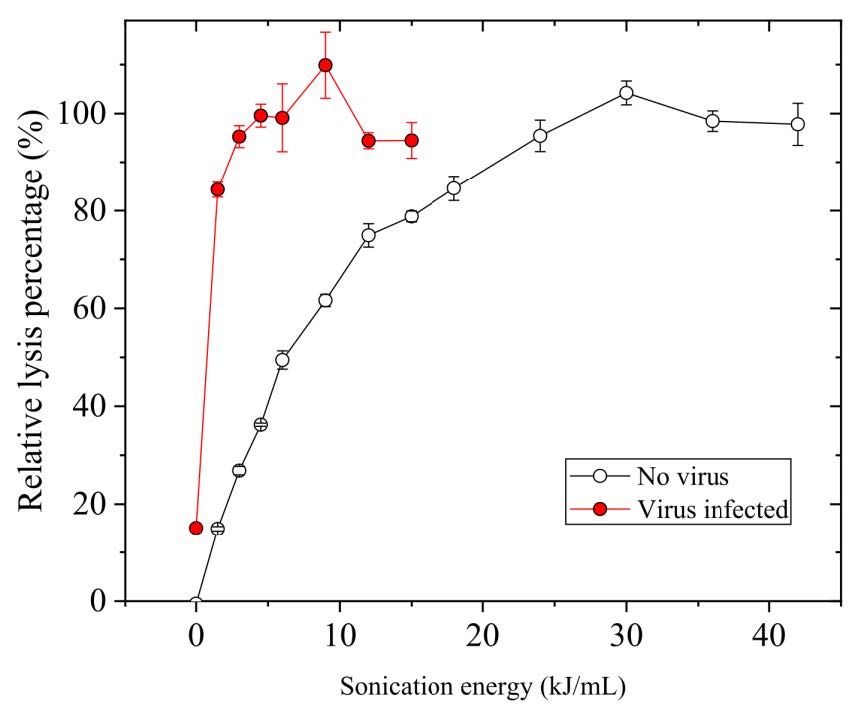
Figure 3. Improvement of cell lysis efficiency after viral infection.
|
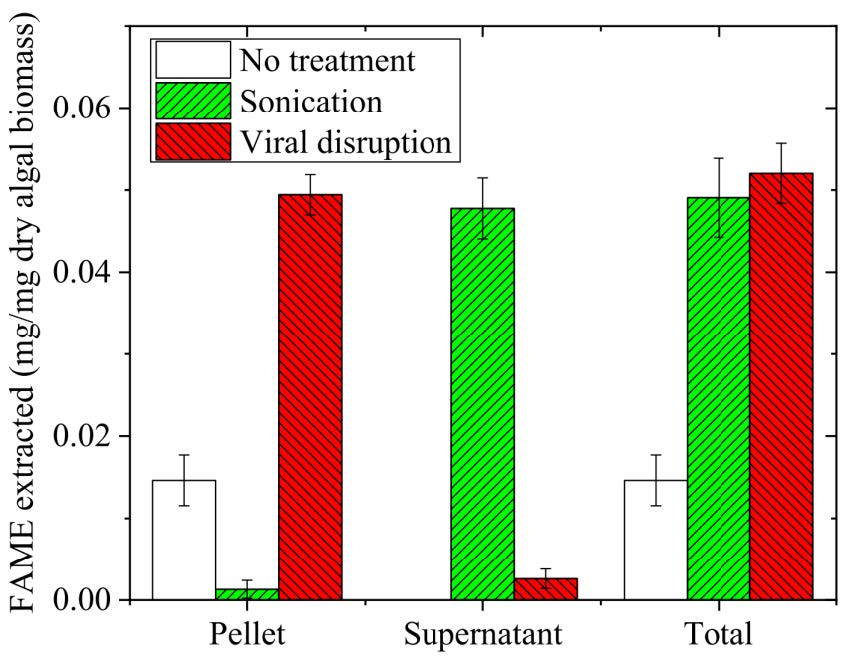
Figure 4. Comparison of FAMEs extracted from algal cells with different disruption methods.
|
 |
 |
|
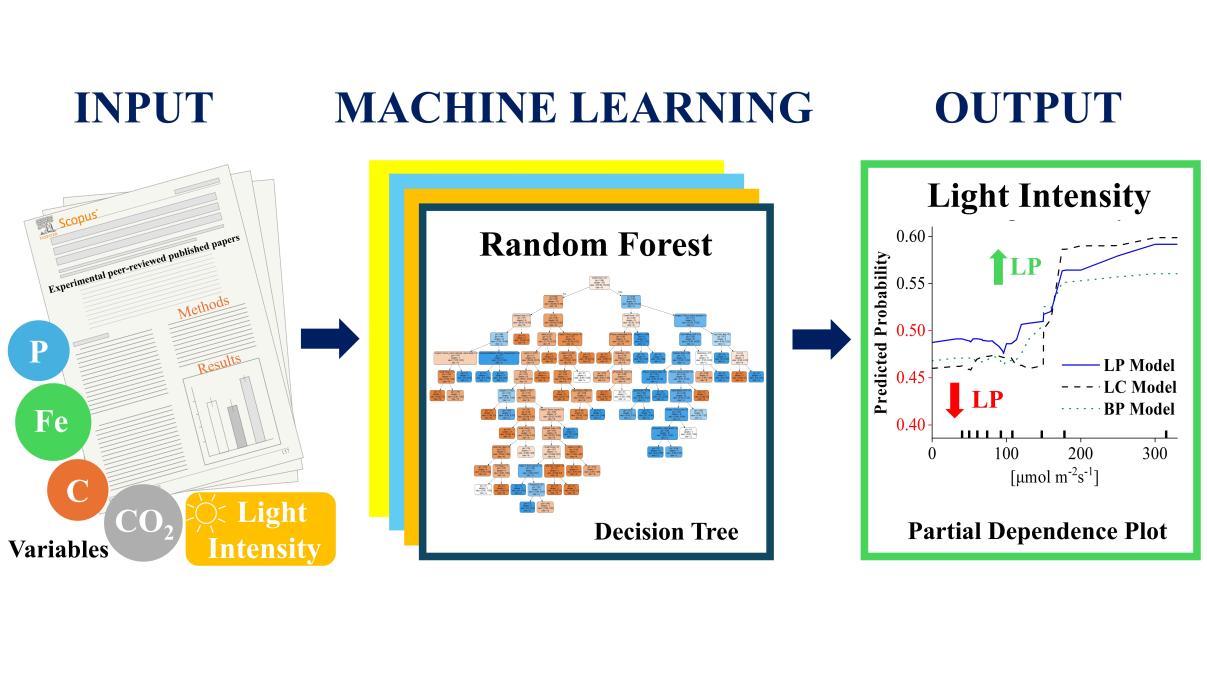
Figure 5. Enhanced biomimetic algal lipid enrichment for improved biofuel production driven by non-stress viral lysis.
|
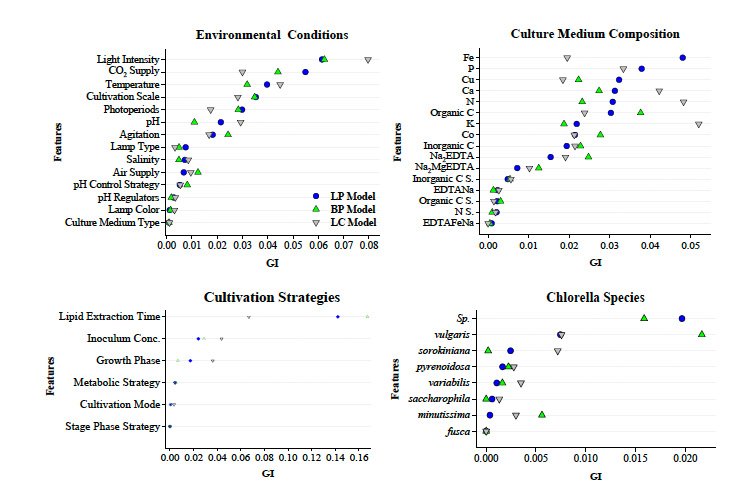
Figure 6. Relative GI rankings for individual cultivation features in the LP, BP, and LC ML-RF models.
|
 |
 |
|
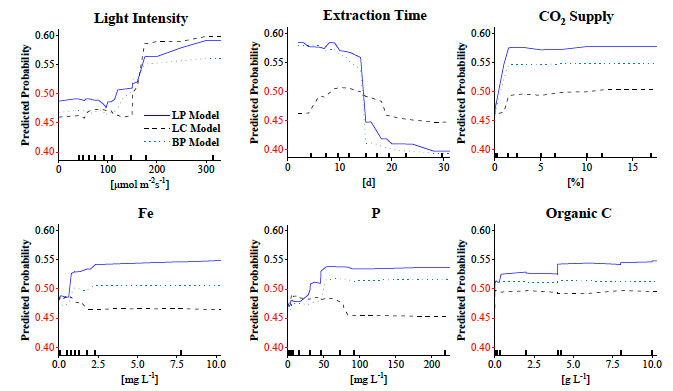
Figure 7. ML-Driven continuous PDPs of the features with the highest likelihood of achieving high LP. Significant operational ranges have absolute probabilities 〉0.5 and do not plateau. Data deciles are denoted with tick marks.
|
Figure 8. Novel machine learning unlocks high lipid productivity and resolves trade-offs in algal biofuel production.
|
 |
 |
|
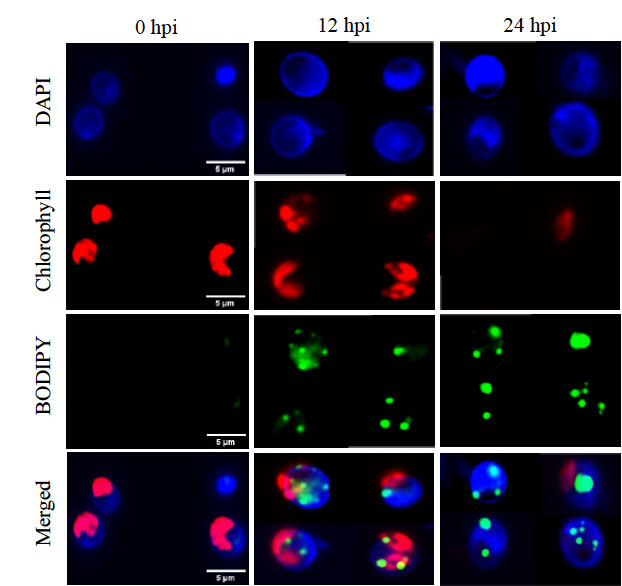
Figure 9. Epifluorescence microscopy images of Chlorella cells infected with PBCV-1, stained with BODIPY 505/515 to visualize neutral lipids (green) and DAPI to highlight nuclear content (blue), as well as intrinsic chlorophyll fluorescence (red). Composite images merging all three fluorescence channels illustrate the spatial overlap and dynamic reorganization of cellular components during viral infection (0,12, and 24 hpi).
|
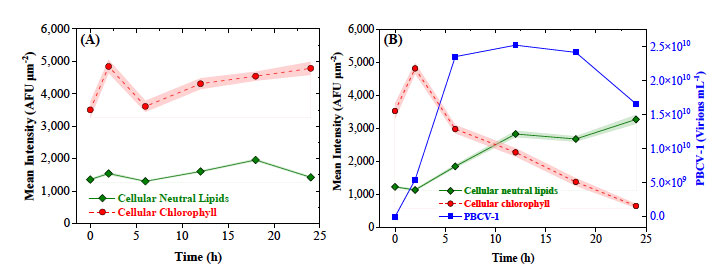
Figure 10. Temporal trajectories of Chlorella’s cellular fluorescent markers (chlorophyll and TAGs) of (A) uninfected control cells and (B) cells infected with PBCV-1 across the full progression of the PBCV-1 lytic cycle.Figure 10. Temporal trajectories of Chlorella’s cellular fluorescent markers (chlorophyll and TAGs) of (A) uninfected control cells and (B) cells infected with PBCV-1 across the full progression of the PBCV-1 lytic cycle.
|
|







































